| Predicted mutation | ||||||
|---|---|---|---|---|---|---|
| evidence | seq id | position | mutation | annotation | gene | description |
| MC JC | NC_000913 | 255,892 | Δ18,063 bp | IS5‑mediated | gpt–[ykfC] | 26 genes gpt, frsA, crl, insB9, insA9, crl, phoE, proB, proA, thrW, ykfI, yafW, ykfH, ykfG, yafX, ykfF, ykfB, yafY, yafZ, ykfA, perR, insN, insI1, insN, eyeA, [ykfC] |
| Missing coverage evidence... | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| seq id | start | end | size | ←reads | reads→ | gene | description | |||
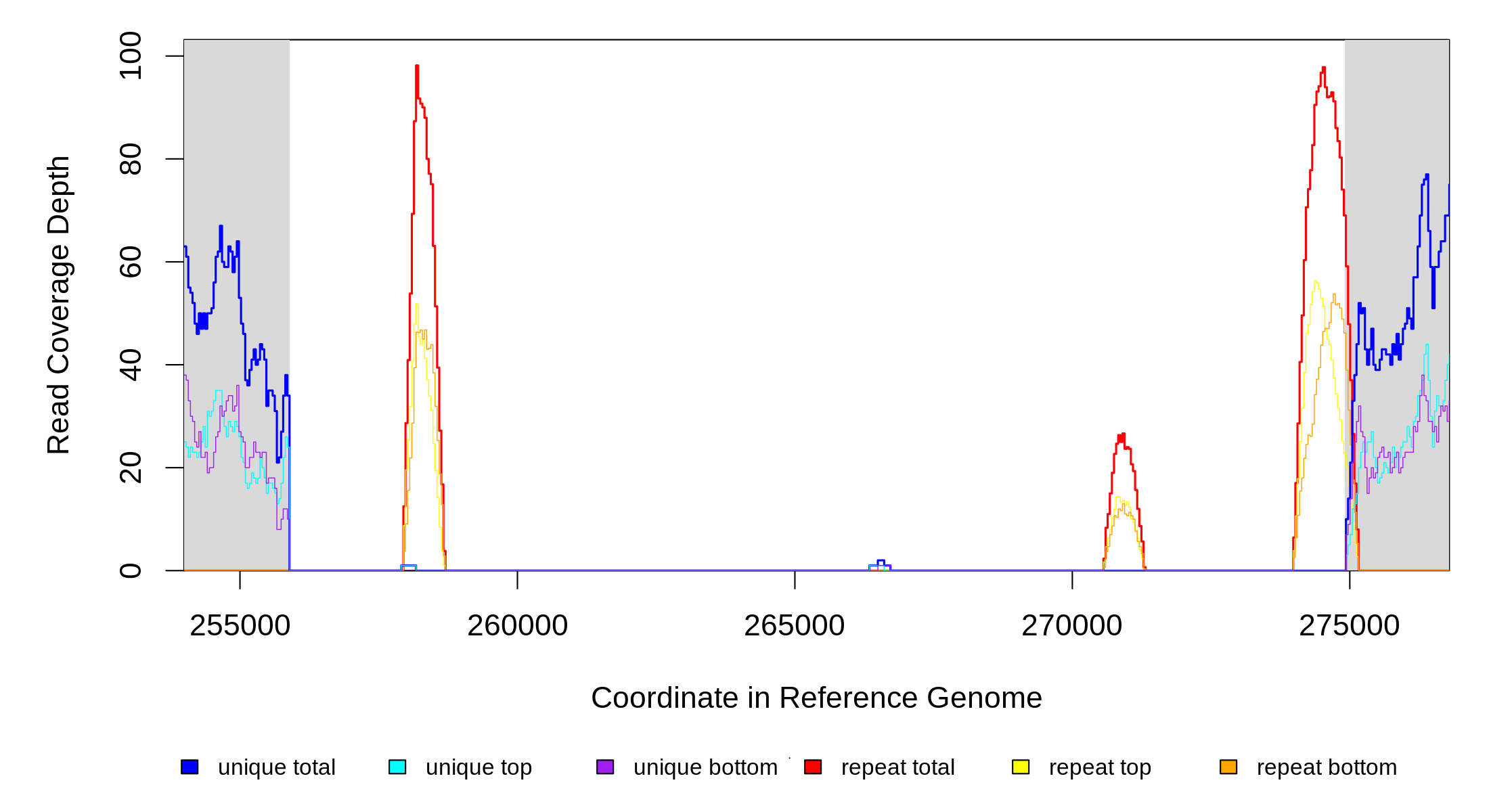
| * | * | ÷ | NC_000913 | 255892 | 274910–273955 | 18064–19019 | 36 [0] | [2] 3 | gpt–[insH1] | gpt,frsA,crl,insB9,insA9,crl,phoE,proB,proA,thrW,ykfI,yafW,ykfH,ykfG,yafX,ykfF,ykfB,yafY,yafZ,ykfA,perR,insN,insI1,insN,eyeA,ykfC,[insH1] |
| New junction evidence | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| seq id | position | reads (cov) | reads (cov) | score | skew | freq | annotation | gene | product | ||
| * | ? | NC_000913 | = 255891 | 0 (0.000) | 36 (0.650) | 36/488 | 0.4 | 100% | intergenic (‑175/‑86) | pepD/gpt | peptidase D/xanthine‑guanine phsophoribosyltransferase |
| ? | NC_000913 | 273955 = | NA (NA) | noncoding (1195/1195 nt) | IS5 | repeat region | |||||
