A very laborious investigation of the distribution of chromosome combinations in a large F2 was carried out by Kihara and Matsumura (1940). When all the disturbing factors were taken into consideration, the frequency of the chromosome combinations obtained from the F2 investigation could be brought into agreement with the results of the other experiments and with the theoretical expectations. Also, the differences between the reciprocal crosses could be explained.
The " sterile " combinations were represented by weak, sterile plants, with the exception of healthy dwarf-like individuals with 20 bivalents and fairly good fertility. At first four different dwarfs were found; in each of them another chromosome pair of the D genome was lacking. Much later Kihara's collaborator, Matsumura (1947), found the three remaining dwarfs. These plants were the first ever produced nullisomics which became so important as instruments in investigations of the gene content of the vulgare chromosomes.
Classification of Aegilops
The genus Aegilops is believed to be originated in Mediterranean districts, namely Asia Minor, Syria and Palestine (Zhukovsky 1928 and Eig 1929), and various species are found naturally inthose districts at the present time.
From the morphological standpoint, Eig classified them into six sections and 22 species, while Zhukovsky into nine sections and 20 species. Considering the results of karyotype analysis by Senianinova-Korczagina (1932), Kihara (1949) has set up six sections, Polyeides, Cylindropyrum, Comopyrum, Amblyopyrum, Sitopsis and Vertebrata. Gastropyrum by Senianinova-Korczagina was included in section Vetebrata by Kihara.
In Aegilops, polyploidy relation with the basic number of seven is found. From the results of genome analyses, 9 different genomes, C, Cu, D, M, Mu, Mt, S, Sb and Sl, have been distinguished among 12 diploid Aegilops specics. Their genome relationships in the Aegilops group, are as follows:
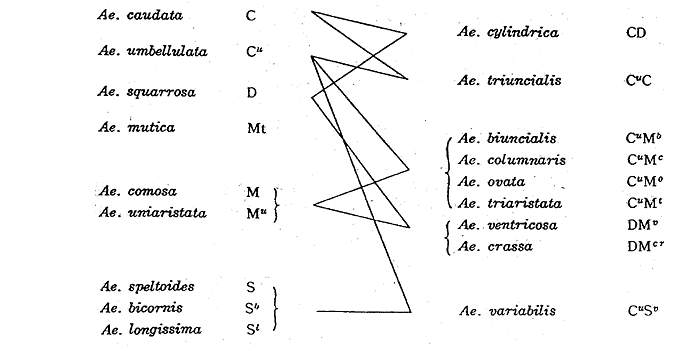
According to genome analyses Ae. Aucheri is related to Ae. speltoides, so Ae. Heldreichii is to Ae. comosa, and so Ae. sharonensis is to Ae. longissima. Ae. Aucheri is different from Ae. speltoides only by one gene for awn. In the hybrids Ae. comosa x Ae. Heldreichii and Ae. longissima x Ae. sharonensis, a ring of 4 chromosomes due to reciprocal translocation is found in addition to 5 pairs. Similarily 2 rings of 4 chromosomes are found in the hybrid, Ae. kotschyi x Ae. variabilis.
In Ae. triaristata and Ae.crassa, tetraploid and hexaploid plants are known. There are 300 possible hybrid combinations in Aegilops, among which 123 combinations have been already raised and cytologically studied.