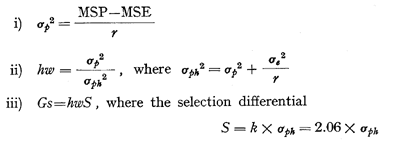Wheat improvement by induced
mutations
A.S. LARIK and H.M.I. HAFIZ*
Department of Plant Breeding and Genetics Sind Agricultural
University, Tandojam, Pakistan
Induced mutations, a method of generating genetic
variability have played a significant role in the evolution
of tribe Triticeae (LARIK 1975a, b; 1976; 1977). In
recent years induced mutations have been successful for the
genetic modifications of many crop plants particularly bread
wheat (Triticum aestivum L.) The philosophy and
achievements of mutation breeding are indeed too well known
(SIGURBJORNSSON and MIKE 1974; BOROJEVIC 1979). Stable
mutants have been isolated from contemporary varieties of
bread wheat with respect to various quantitative traits
(LARIK 1978a, b) including disease resistance (SIDDIQUI and
SIDDIQUI 1974), lodging resistance (LARIK et al.
1980a) and improved protein content and quality (SIDDIQUI
et al. 1975).
Our main objective of wheat improvement however, is the
induction and accumulation of positive variation associated
with the advancement of grain yield and its effective
components. Phenotypically stable wheat mutants were
compared with a commercial variety (Pak-70) which covers the
major wheat growing areas in Pakistan. However, this variety
is susceptible to rusts and is showing considerable genetic
deterioration with the passage of time. The present paper
discusses the evolution of different mutants which are
superior to existing commercial variety in grain yield and
other useful agronomic traits.
Materials and Methods
Phenotypically stable mutants of three hexaploid wheat
cultivars viz., Pak-70, Nayab and 6134 x C-271 were
evaluated in M6 generation and compared with a
commercial variety during Rabi 1979-80 at Tandojam, Sind,
Pakistan. Seeds of these mutants and mother cultivars were
planted in 5 rows, 3 m long with 30 cm distance between rows
in a randomized complete block design with 4
replications.
Analysis of variance for yield and other metrical traits was
carried out separately. The pertinent mean squares and
parameters estimated in each analysis were as follows:
|
Source of variation |
Mean squares |
Mean square expectations |
|
Genotypes (Strains) |
MSP |
sigmae2+rsigmap2 |
|
Error |
MSE |
sigmae2 |
where sigmae2, is the error variance, sigmap2 is the component variance due to genetic difference among strains and r is the number of replications. The selection parameters such as genotypic variance among strains (sigmap2), heritability (hw) and genetic gain expected from selection (Gs) were determined similar to GHAFOOR ARAIN (1973) as under: